Flux Spectrum and E-Index Analysis
This example demonstrates how to attach a neutron flux spectrum to a Benchmark using a Serpent detector file, visualize it with FluxSpectrum.plot_spectrum, and compute the E-index similarity matrix via AssimilationSuite.e_index_matrix. The E-index quantifies the cosine similarity between sensitivity profiles without the influence of a nuclear data covariance matrix.
1. Setup and Imports
We import andalus for the core logic and matplotlib for visualization.
[1]:
import matplotlib.pyplot as plt
import andalus
2. Reading Serpent Output
We load the HMF001 (Godiva) benchmark using Benchmark.from_serpent. The optional flux_det argument attaches a FluxSpectrum by reading a named detector from a Serpent _det0.m file. Here we pass the detector name "flux" and the path to the detector output file.
The pertlist filters the sensitivity output to the most physically relevant reactions:
mt 2 xs: Elastic scatteringmt 18 xs: Fissionmt 102 xs: Radiative capturenubar prompt: Average prompt neutron multiplicitychi prompt: Prompt fission neutron spectrum
[2]:
hmf001 = andalus.Benchmark.from_serpent(
title="HMF001",
m=1.0,
dm=0.001,
sens0_path="../data/hmf001.ser_sens0.m",
results_path="../data/hmf001.ser_res.m",
pertlist=["mt 2 xs", "mt 18 xs", "mt 102 xs", "nubar prompt", "chi prompt"],
flux_det=("flux", "../data/hmf001.ser_det0.m"),
)
Reading ../data/hmf001.ser_res.m
WARNING: SERPENT Serpent 2.2.1 found in ../data/hmf001.ser_res.m, but version 2.1.31 is defined in settings
WARNING: Attempting to read anyway. Please report strange behaviors/failures to developers.
- done
Reading ../data/hmf001.ser_sens0.m
- done
Reading ../data/hmf001.ser_det0.m
- done
The flux attribute is a FluxSpectrum, a pandas DataFrame subclass with a two-level (E_min_eV, E_max_eV) MultiIndex. We inspect the first few rows to verify the energy grid and flux values.
[3]:
hmf001.flux.head()
[3]:
| flux | flux_std | ||
|---|---|---|---|
| E_min_eV | E_max_eV | ||
| 0.00001 | 0.10000 | 0.000000e+00 | 0.000000e+00 |
| 0.10000 | 0.54000 | 0.000000e+00 | 0.000000e+00 |
| 0.54000 | 4.00000 | 0.000000e+00 | 0.000000e+00 |
| 4.00000 | 8.31529 | 5.409640e-09 | 5.409640e-09 |
| 8.31529 | 13.70960 | 0.000000e+00 | 0.000000e+00 |
3. Flux Spectrum Characterization
Two scalar metrics summarize where neutrons are most active in energy:
where \(\bar{E}_i = \sqrt{E_{\min,i} \cdot E_{\max,i}}\) is the geometric mean energy of bin \(i\). EALF is the flux-weighted geometric mean (sensitive to the lethargy distribution), while \(\langle E \rangle\) is the flux-weighted arithmetic mean. For a fast spectrum such as HMF001 we expect both to lie well above the thermal region.
[4]:
print(f"EALF: {hmf001.flux.ealf:.3e} eV")
print(f"Mean energy: {hmf001.flux.mean_energy:.3e} eV")
EALF: 8.474e+05 eV
Mean energy: 1.461e+06 eV
4. Plotting the Flux Spectrum
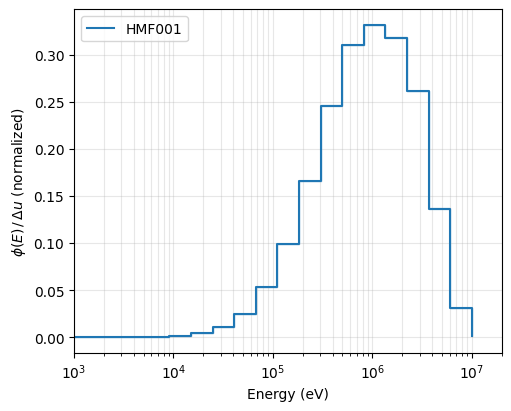
We plot the normalized flux per unit lethargy, \(\phi(E)/\Delta u\), on a logarithmic energy axis. The shaded band represents the \(1\sigma\) statistical uncertainty from the Serpent tally. The sharp peak in the fast region is characteristic of the bare HEU Godiva configuration.
[9]:
fig, ax = plt.subplots(figsize=(5, 4), layout="constrained")
hmf001.flux.plot_spectrum(ax=ax)
ax.set(xlim=(1e3, 2e7))
plt.show()
5. E-Index Similarity via AssimilationSuite
The E-index measures the cosine similarity between two sensitivity vectors:
Unlike \(c_k\), the E-index does not involve a covariance matrix, so it captures purely geometric similarity of the sensitivity profiles. We construct a minimal AssimilationSuite with HMF001 as both the benchmark and the application to illustrate the API.
[7]:
suite = andalus.AssimilationSuite.from_yaml("config.yaml")
Reading ../data/hmf001.ser_res.m
WARNING: SERPENT Serpent 2.2.1 found in ../data/hmf001.ser_res.m, but version 2.1.31 is defined in settings
WARNING: Attempting to read anyway. Please report strange behaviors/failures to developers.
- done
Reading ../data/hmf001.ser_sens0.m
- done
Reading ../data/hmf002-001.ser_res.m
WARNING: SERPENT Serpent 2.2.1 found in ../data/hmf002-001.ser_res.m, but version 2.1.31 is defined in settings
WARNING: Attempting to read anyway. Please report strange behaviors/failures to developers.
- done
Reading ../data/hmf002-001.ser_sens0.m
- done
Reading ../data/hmf002-002.ser_res.m
WARNING: SERPENT Serpent 2.2.1 found in ../data/hmf002-002.ser_res.m, but version 2.1.31 is defined in settings
WARNING: Attempting to read anyway. Please report strange behaviors/failures to developers.
- done
Reading ../data/hmf002-002.ser_sens0.m
- done
[8]:
e_index = suite.e_index_matrix()
print(e_index)
HMF001 HMF002-001 HMF002-002
HMF001 1.000000 0.960530 0.957067
HMF002-001 0.960530 1.000000 0.999763
HMF002-002 0.957067 0.999763 1.000000
The diagonal entries equal 1.0 by construction. The off-diagonal value between the benchmark and application reflects how closely aligned their sensitivity profiles are in the multivariate nuclear-data space. An E-index near 1.0 indicates that the two systems are driven by the same energy and reaction dependencies, making the benchmark a strong predictor of the application’s behavior.