Processing ERRORR Files with Andalus
This notebook demonstrates how to create an andalus.Covariance object from ERRORR output files and prepare the data for database storage.
Background
We use the sandy library to parse ERRORR files and combine covariance matrices. Currently, andalus supports reading covariance information from the following sections:
File 31 (MF31): \(\bar{\nu}\) (nubar)
File 33 (MF33): Cross sections
File 34 (MF34): Angular distributions
File 35 (MF35): Energy distributions
Note on Identifiers: Since MT numbers in MF34 and MF35 may overlap with those in MF33, we ensure uniqueness in MF34/35 by defining the identifier as:Identifier = (MF * 1000) + MT
1. Setup and Imports
We import andalus to handle the covariance objects. Internally, andalus leverages sandy to parse the raw text into structured pandas DataFrames.
[126]:
import matplotlib.pyplot as plt
import andalus
2. Reading ERRORR Output
Raw ERRORR files use a specific nuclear data format that is difficult to parse manually. Below is a snippet of the raw input data:
[37]:
!head -10 "../data/u235.errorr31"
1 0 0 0
9.223500+4 2.330248+2 6 0 -11 09228 1451 1
0.000000+0 0.000000+0 33 0 34 09228 1451 2
1.000000-5 1.000000-1 5.400000-1 4.000000+0 8.315290+0 1.370960+19228 1451 3
2.260330+1 4.016900+1 6.790400+1 9.166090+1 1.486250+2 3.043250+29228 1451 4
4.539990+2 7.485180+2 1.234100+3 2.034680+3 3.354630+3 5.530840+39228 1451 5
9.118820+3 1.503440+4 2.478750+4 4.086770+4 6.737950+4 1.110900+59228 1451 6
1.831560+5 3.019740+5 4.978710+5 8.208500+5 1.353350+6 2.231300+69228 1451 7
3.678790+6 6.065310+6 1.000000+7 1.964030+7 9228 1451 8
9228 1 099999
The Covariance.from_errorr method simplifies this process. You provide a dictionary mapping the error types (keys) to their file paths, specify the ZAI (isotope identifier), and optionally provide a list of MT numbers to extract.
[133]:
files = {
"errorr33": "../data/u235.errorr33",
"errorr31": "../data/u235.errorr31",
"errorr34": "../data/u235.errorr34",
"errorr35": "../data/u235.errorr35",
}
cov = andalus.Covariance.from_errorr(files, zai=922350, mts=[1, 2, 4, 18, 102, 456, 34251, 35018])
3. Exploring the Covariance Object
The resulting object inherits from pandas.DataFrame, meaning you can use standard data manipulation tools. However, it also contains nuclear-specific metadata like zai, nuclide, and optional physics parameters like temperature or tolerance.
[134]:
print(f"ZAI: {cov.zai}")
print(f"Nuclide: {cov.nuclide}")
?cov
ZAI: 922350
Nuclide: U235
Type: Covariance
String form:
MT 1 \
E_min_eV <...> 1.00000e+07 3.95538e-03
1.00000e+07 1.96403e+07 1.96065e-02
[264 rows x 264 columns]
Length: 264
File: c:\users\dhouben\documents\andalus\src\andalus\covariance.py
Docstring:
Subclass of pd.DataFrame for handling individual nuclide covariance data.
Attributes
----------
zai : int
Nuclide identifier (ZAI).
temperature : float, optional
Temperature at which the covariance was generated.
err : float, optional
Associated error or tolerance.
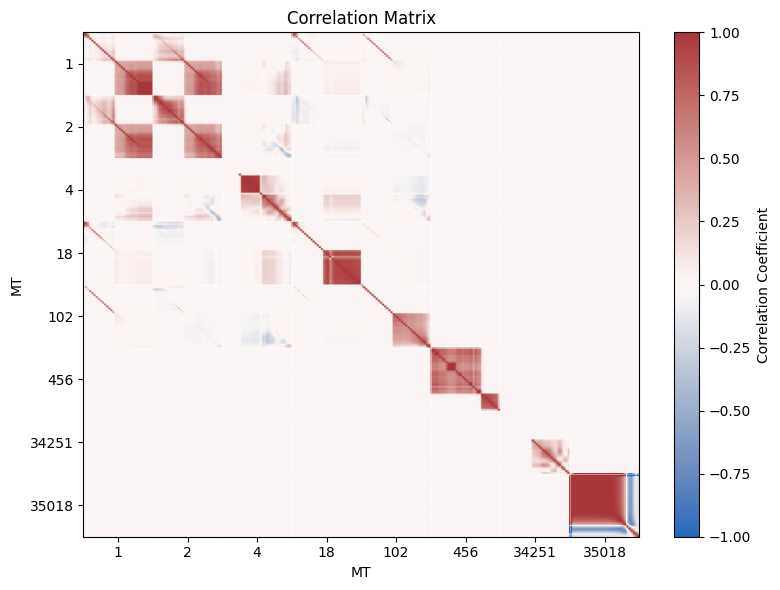
4. Correlation Matrix Visualization
We can easily derive a correlation matrix from our covariance data and visualize it using a heat map.
[135]:
corr = cov.correlation()
fig, ax = plt.subplots(figsize=(8, 6))
im = ax.imshow(corr.values, vmin=-1, vmax=1, cmap="vlag", aspect="auto")
cbar = fig.colorbar(im, ax=ax)
cbar.set_label("Correlation Coefficient")
ax.set_title("Correlation Matrix")
ax.set_xlabel("Index")
ax.set_ylabel("Index")
lvl0 = corr.index.get_level_values(0)
# find contiguous MT blocks
blocks = []
start = 0
for i in range(1, len(lvl0)):
if lvl0[i] != lvl0[i - 1]:
blocks.append((lvl0[i - 1], start, i - 1))
start = i
blocks.append((lvl0[-1], start, len(lvl0) - 1))
# tick at block centers
tick_pos = [(s + e) / 2 for _, s, e in blocks]
tick_labels = [str(mt) for mt, _, _ in blocks]
ax.set_xticks(tick_pos)
ax.set_xticklabels(tick_labels)
ax.set_yticks(tick_pos)
ax.set_yticklabels(tick_labels)
# optional separators between MT blocks
for _, _, e in blocks[:-1]:
ax.axvline(e + 0.5, color="white", lw=0.5, alpha=0.6)
ax.axhline(e + 0.5, color="white", lw=0.5, alpha=0.6)
ax.set_xlabel("MT")
ax.set_ylabel("MT")
plt.tight_layout()
5. Exporting Data
Once the data is processed, it can be exported to a format optimized for use in ANDALUS.
[53]:
cov.to_hdf5("u235_cov.h5")
And it can then be easily read.
[66]:
andalus.Covariance.from_hdf5("u235_cov.h5", zai=922350)
[66]:
| MT | 456 | ... | 35018 | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E_min | 1.00000e-05 | 1.00000e-01 | 5.40000e-01 | 4.00000e+00 | 8.31529e+00 | 1.37096e+01 | 2.26033e+01 | 4.01690e+01 | 6.79040e+01 | 9.16609e+01 | ... | 1.11090e+05 | 1.83156e+05 | 3.01974e+05 | 4.97871e+05 | 8.20850e+05 | 1.35335e+06 | 2.23130e+06 | 3.67879e+06 | 6.06531e+06 | 1.00000e+07 | ||
| E_max | 1.00000e-01 | 5.40000e-01 | 4.00000e+00 | 8.31529e+00 | 1.37096e+01 | 2.26033e+01 | 4.01690e+01 | 6.79040e+01 | 9.16609e+01 | 1.48625e+02 | ... | 1.83156e+05 | 3.01974e+05 | 4.97871e+05 | 8.20850e+05 | 1.35335e+06 | 2.23130e+06 | 3.67879e+06 | 6.06531e+06 | 1.00000e+07 | 1.96403e+07 | ||
| MT | E_min | E_max | |||||||||||||||||||||
| 456 | 1.00000e-05 | 1.00000e-01 | 2.05760e-05 | 2.05760e-05 | 1.79154e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | ... | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 |
| 1.00000e-01 | 5.40000e-01 | 2.05760e-05 | 2.05760e-05 | 1.79154e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | ... | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | |
| 5.40000e-01 | 4.00000e+00 | 1.79154e-05 | 1.79154e-05 | 2.00769e-05 | 2.03007e-05 | 1.84709e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | ... | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | |
| 4.00000e+00 | 8.31529e+00 | 1.76400e-05 | 1.76400e-05 | 2.03007e-05 | 2.05760e-05 | 1.85570e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | ... | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | |
| 8.31529e+00 | 1.37096e+01 | 1.76400e-05 | 1.76400e-05 | 1.84709e-05 | 1.85570e-05 | 2.71709e-05 | 1.95905e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | 1.76400e-05 | ... | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 35018 | 1.35335e+06 | 2.23130e+06 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | ... | -8.24541e-04 | -6.39354e-04 | -3.54139e-04 | -1.32770e-04 | 4.91341e-05 | 2.42313e-04 | 1.20013e-04 | 4.55191e-06 | -9.22334e-05 | -2.44922e-06 |
| 2.23130e+06 | 3.67879e+06 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | ... | -7.69174e-04 | -6.44898e-04 | -4.72111e-04 | -2.48225e-04 | -3.08282e-05 | 1.20013e-04 | 2.92605e-04 | 2.06564e-04 | 4.78707e-05 | -3.31639e-06 | |
| 3.67879e+06 | 6.06531e+06 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | ... | -6.33792e-04 | -5.84774e-04 | -4.77911e-04 | -3.00563e-04 | -8.00144e-05 | 4.55191e-06 | 2.06564e-04 | 5.42362e-04 | 6.37521e-04 | 7.84379e-04 | |
| 6.06531e+06 | 1.00000e+07 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | ... | -5.45683e-04 | -5.36010e-04 | -4.44288e-04 | -2.73286e-04 | -1.02590e-04 | -9.22334e-05 | 4.78707e-05 | 6.37521e-04 | 2.10911e-03 | 3.95538e-03 | |
| 1.00000e+07 | 1.96403e+07 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | 0.00000e+00 | ... | -1.32210e-03 | -8.83098e-04 | -4.94075e-04 | -3.63020e-04 | -2.68487e-04 | -2.44922e-06 | -3.31639e-06 | 7.84379e-04 | 3.95538e-03 | 1.96065e-02 | |
231 rows × 231 columns